Parameterization
The Poisson distribution for a random vector \(\pmb{y} = (y_1, y_2, \dots, y_n)\) is defined by the probability mass function:
\[f(y_i \mid \lambda_i) = \frac{\lambda_i^{y_i} e^{-\lambda_i}}{y_i!}, \quad y_i = 0, 1, 2, \dots, \quad i = 1, 2, \dots, n\]
where \(\pmb{\lambda} = (\lambda_1,
\lambda_2, \dots, \lambda_n)\) is a vector of rate parameters,
with \(\lambda_i > 0\) representing
the mean number of events for each observation \(y_i\).
Assume \(n\) = 1. To illustrate how the
Poisson PMF changes for different values of the rate parameter \(\lambda\), Figure 1 displays two subplots. The left
subplot shows \(\lambda\) in \(\{1,2,3\}\), while the right subplot covers
larger rates (\(\lambda \in
\{5,10,15\}\)).
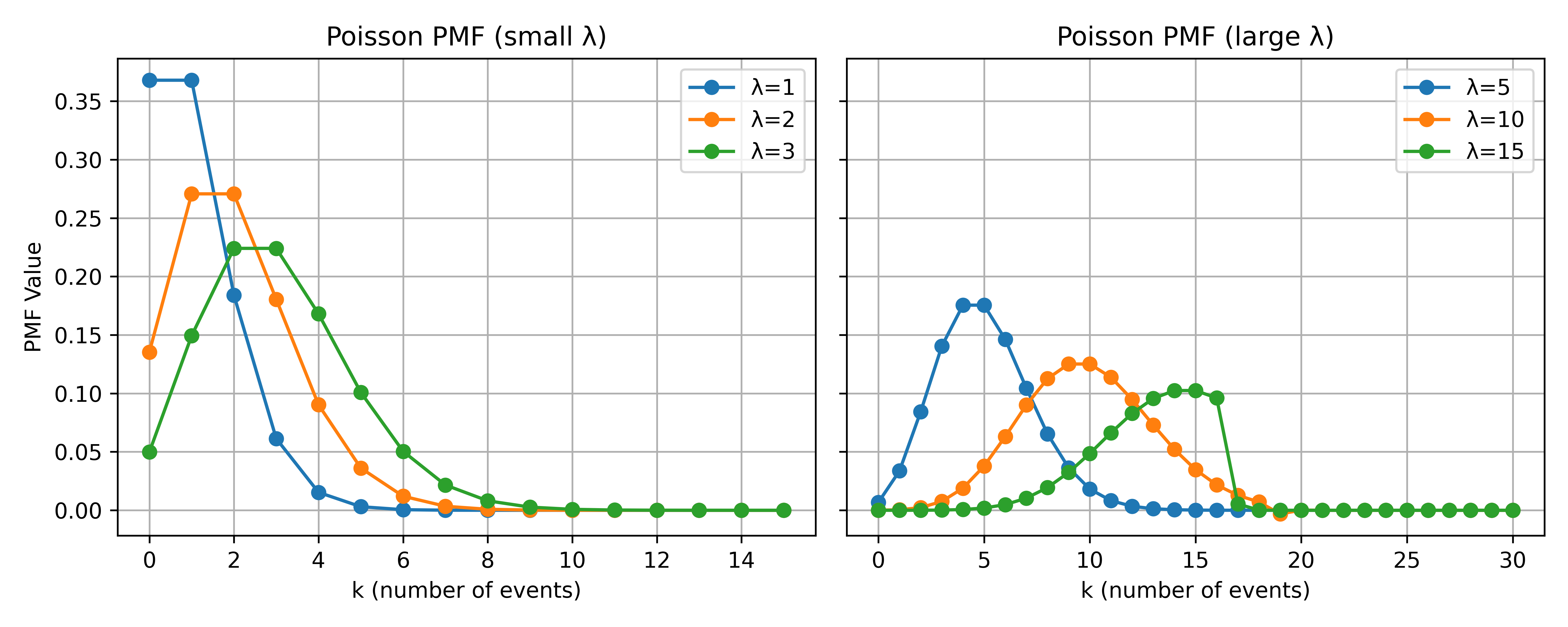
For smaller values of \(\lambda\), the distribution is more concentrated around lower counts. As \(\lambda\) grows, the distribution shifts rightward and spreads out, reflecting the higher expected number of events.
Link Function
The canonical link function for the Poisson distribution is the log link:
\[\eta_i = \log(\lambda_i), \quad i = 1, 2, \dots, n\]
The relationship between the linear predictor \(\pmb{\eta} = (\eta_1, \eta_2, \dots, \eta_n)\) and the mean \(\pmb{\lambda}\) is given by:
\[\lambda_i = \exp(\eta_i), \quad i = 1, 2, \dots, n\]
or in vector form:
\[\pmb{\lambda} = \exp(\pmb{\eta})\]
Hyperparameters
The Poisson likelihood has no hyperparameters. The rate parameter \(\lambda\) is fully determined by the linear predictor \(\eta\) through the log link function.
Validation Rules
pyINLA enforces several validation rules for Poisson models to ensure correct specification:
No Hyper Configuration
The Poisson likelihood has no hyperparameters. Do NOT provide control['family']['hyper'] configuration.
Exposure (E)
Key: E
When providing the E (exposure) argument:
All values must be strictly positive (> 0)
Length must match the number of observations
# With exposure
result = pyinla(model={'response': 'y', 'fixed': ['1', 'x']}, data=df, family="poisson", E=df["exposure"])
Offset Alternative
Key: offset
Instead of E, you can use offset = log(E). When using offset:
Values must not contain Inf; NaN is allowed and treated as 0 (no offset for that observation)
Length must match the number of observations
import numpy as np
# Using offset instead of E
result = pyinla(model={'response': 'y', 'fixed': ['1', 'x']}, data=df, family="poisson",
offset=np.log(df["exposure"]))
Response Values
Response variable \(\pmb{y}\) must be non-negative integers (counts: 0, 1, 2, ...).
Allowed Link Functions
Key: control['family']['link']
These link functions are supported:
log(default)logoffsetquantile
Specification
family="poisson"Required arguments:
\(\pmb y\) (integer-valued counts)
Optional arguments:
E: exposure vector (positive values)offset: log-exposure offset