Trainings
Learn Bayesian spatial statistics and pyINLA through structured self-study modules, or book a live one-week intensive course led by the pyINLA team. Tailored for researchers, students, and data teams.
Introduction to Bayesian Inference
Learn the fundamentals: Bayes' theorem, conjugate priors, MCMC sampling, and why INLA is fast. Fit your first Poisson model with pyINLA using the Game of Thrones counting example.
- Bayes' theorem and posterior distributions
- Conjugate analysis (Poisson-Gamma)
- Metropolis-Hastings MCMC
- pyINLA model fitting and posterior transformation
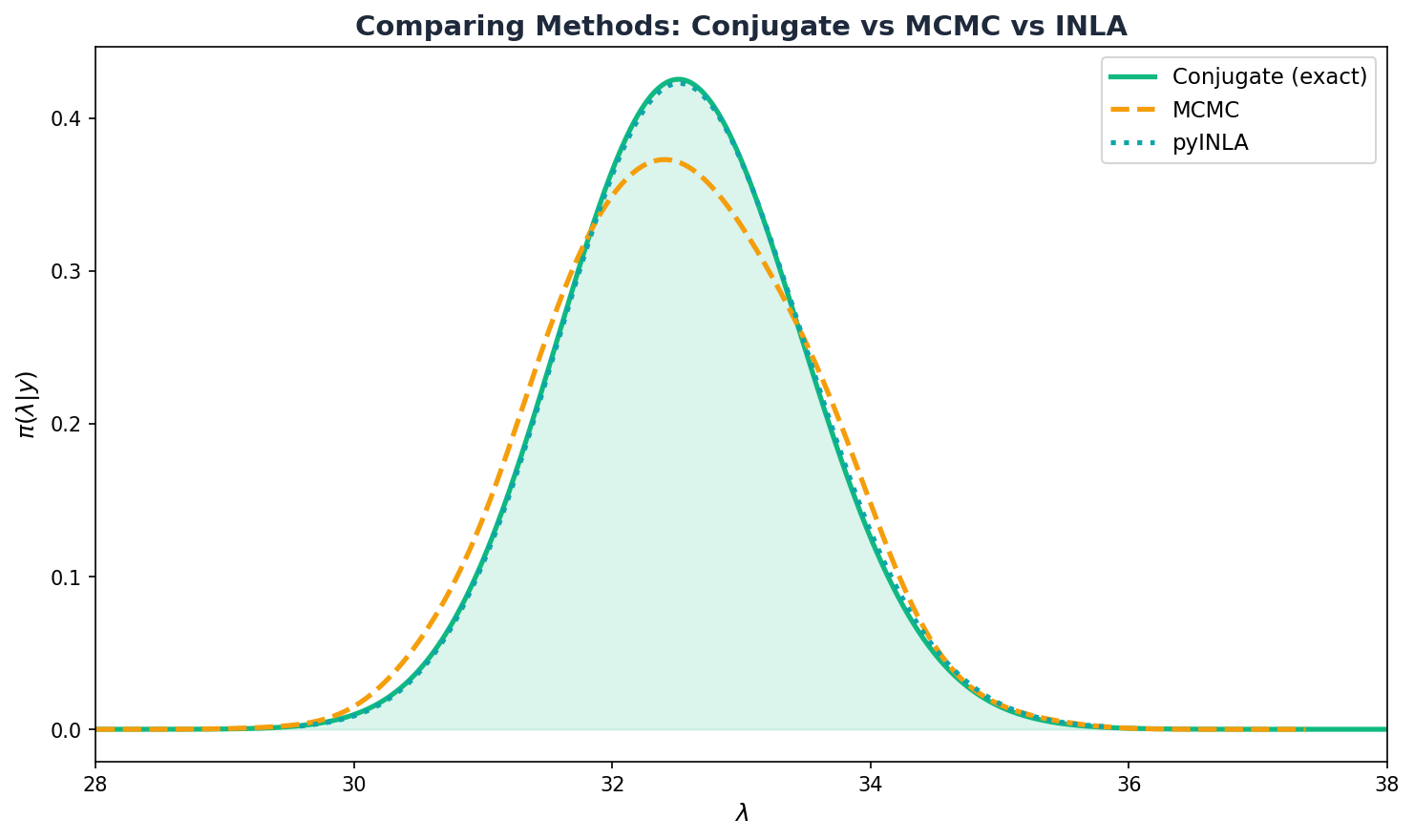
Conjugate vs MCMC vs pyINLA
Regression and Generalized Linear Models
Fit linear regression and Poisson GLMs, compare models with DIC/WAIC/CPO, and work with posterior marginals and sampling.
- Linear regression with pyINLA
- Poisson GLM with offsets
- Model comparison: DIC, WAIC, CPO
- Marginal utilities and posterior sampling
Mixed Effects Models
Add random effects to your models: IID for overdispersion, time series (RW1, AR1), constraints, and generic structures.
- IID random effects for overdispersion
- Time series: RW1 and AR1 models
- Constraints on random effects
- Generic models: Z and custom precision
Multilevel Models
Students in classrooms, patients in hospitals: real data has layers. Learn to model nested and crossed random effects.
- Crossed effects: Penicillin experiment
- Longitudinal data: Sleep deprivation study
- Three-level nesting: Lab quality control
- Logistic multilevel: Election 1988
- Poisson multilevel and overdispersion: NYC Stops
Priors
Choose and customize prior distributions: built-in priors, expression-based custom priors, Penalized Complexity (PC) priors, sensitivity analysis, and model scaling.
- Built-in priors: loggamma, Gaussian, flat, PC
- Custom priors with
expression:syntax - Penalized Complexity (PC) priors
- Sensitivity analysis across prior choices
- Scaling intrinsic random effects (RW1, RW2, Besag)
Spatial Models
Areal data, geostatistics, and point patterns: model spatial autocorrelation with lattice models, SPDE, and adjacency structures.
- Regular lattice: RW2D and Matern2D models
- Irregular lattice: Besag, BYM, and Leroux models
- Geostatistics: SPDE with Matern covariance
- Point patterns: Log-Gaussian Cox processes
- Model comparison across spatial specifications
Temporal Models
Model time series and spatio-temporal data: AR(1), RW1, forecasting with NAs, and Kronecker products for space-time interaction.
- AR(1) and RW1 for climate reconstruction
- Poisson time series: earthquake counts
- Forecasting with the NA trick
- Spatio-temporal: Besag x AR1 Kronecker models
- Informative priors and model comparison
Smoothing
Estimate flexible, data-driven curves with penalized splines: RW1/RW2 random walks, binned covariates, and 1D SPDE for non-parametric regression.
- Polynomial regression as a baseline
- Penalized splines: RW1 and RW2 models
- Binned covariates for computational efficiency
- Non-Gaussian smoothing: binomial dose-response
- 1D SPDE with Matern covariance
Survival Models
Model censored time-to-event data: exponential, Weibull, lognormal, Cox proportional hazards, and frailty models for clustered survival.
- Exponential, Weibull, and lognormal survival
- Cox proportional hazards with baseline hazard
- Accelerated failure time interpretation
- Frailty models: random effects for clustered data
- Model comparison across survival specifications
Advanced Features
Unlock the full power of pyINLA: predictor matrices, linear combinations, multiple likelihoods, shared terms (copy and replicate), and linear constraints.
- Predictor matrices (A-matrix) for custom designs
- Linear combinations of latent effects
- Multiple likelihoods: joint Gaussian + Poisson models
- Copy and replicate for shared terms
- Linear constraints (sum-to-zero and beyond)
Missing Data
Handle missing observations: predict missing responses automatically, impute missing covariates with plug-in values, and propagate uncertainty with multiple imputation and INLA within MCMC.
- Missingness mechanisms: MCAR, MAR, MNAR
- Predicting missing responses (the NA trick)
- Plug-in imputation for missing covariates
- Multiple imputation with Rubin's rules
- INLA within MCMC for complex imputation
Mixture Models
Fit finite mixture models: handle known groups with multiple likelihoods, estimate unknown group assignments with INLA within MCMC, select the number of components, and fit cure rate survival models.
- Gaussian mixtures with known group assignments
- INLA within MCMC for unknown allocations
- Model selection via marginal likelihoods
- Cure rate models (weibullcure family)
- Label switching and identifiability
Zero-inflated and Hurdle Models
Handle excess zeros common in count data: zero-inflated Poisson and negative binomial models, hurdle models, and strategies for choosing among them.
- Zero-inflation vs. overdispersion: diagnosis and remedies
- Zero-inflated Poisson (ZIP) and ZINB models
- Hurdle Poisson and hurdle negative binomial
- Two-part models with different covariates per component
- Model comparison via DIC, WAIC, and marginal likelihood
Joint Models
Simultaneously model a longitudinal biomarker trajectory and a time-to-event outcome, linked through shared latent effects. Accounts for informative dropout and measurement error.
- Motivating example: CD4 counts and AIDS progression
- Longitudinal sub-model: linear mixed effects with INLA
- Survival sub-model: Cox hazard with shared random effect
- Association structures: current value, current slope, cumulative
- Dynamic predictions and discrimination metrics
Further Reading
Some datasets used in these modules are drawn from the following references. Content, code, and explanations are developed for pyINLA in Python, and go beyond what is covered in these books.
- Gomez-Rubio, V. (2020). Bayesian Inference with INLA. CRC Press. Free online
- Moraga, P. (2019). Geospatial Health Data: Modeling and Visualization with R-INLA and Shiny. CRC Press. Free online
- Moraga, P. (2023). Spatial Statistics for Data Science: Theory and Practice with R. Chapman & Hall/CRC. Free online
- Blangiardo, M., and Cameletti, M. (2015). Spatial and Spatio-temporal Bayesian Models with R-INLA. Wiley.
- Wang, X., Yue, Y.R., and Faraway, J.J. (2018). Bayesian Regression Modeling with INLA. Chapman & Hall/CRC.
- Krainski, E.T., Gómez-Rubio, V., Bakka, H., Lenzi, A., Castro-Camilo, D., Simpson, D., Lindgren, F., and Rue, H. (2019). Advanced Spatial Modeling with Stochastic Partial Differential Equations Using R and INLA. Chapman & Hall/CRC. Free online
- Zuur, A.F., Ieno, E.N., and Saveliev, A.A. (2017). Beginner's Guide to Spatial, Temporal and Spatial-Temporal Ecological Data Analysis with R-INLA. Highland Statistics.