Parametrization
The Negative Binomial distribution for a random vector \(\pmb{y} = (y_1, y_2, \dots, y_m)\) of count observations is defined by the probability mass function:
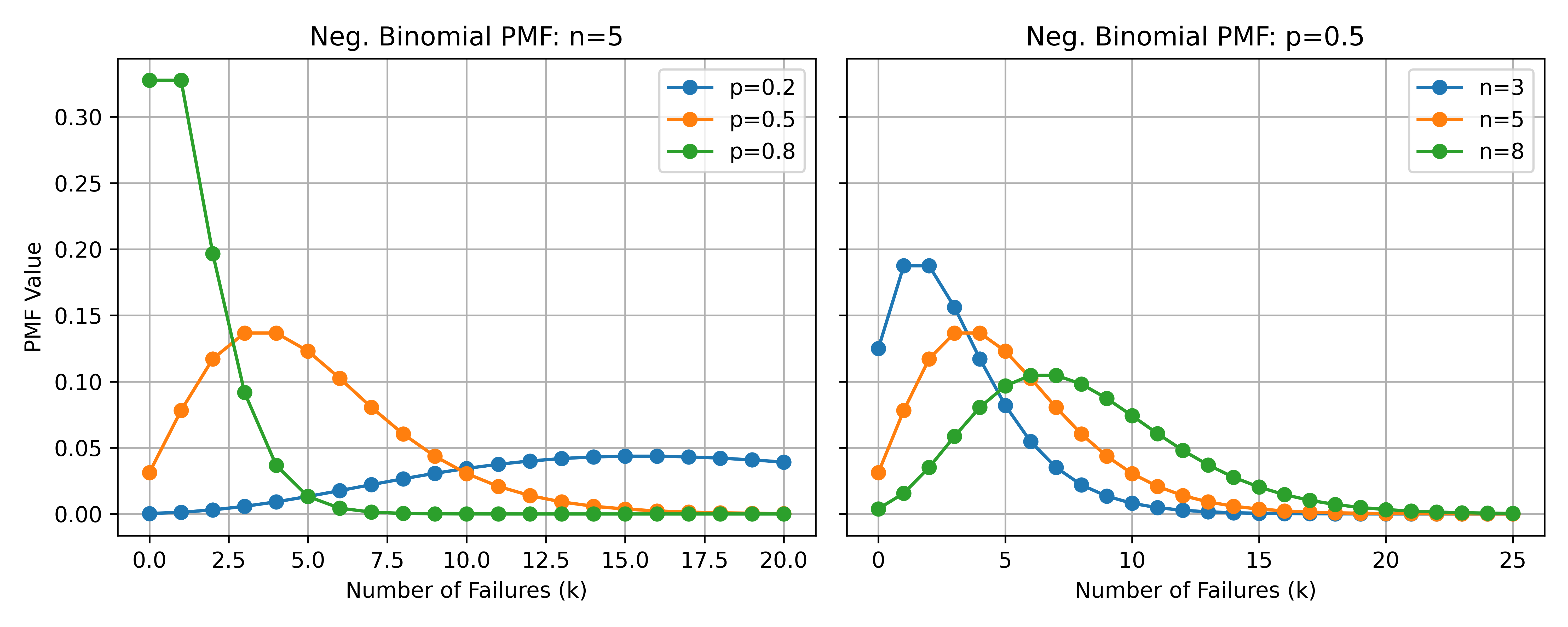
\[\text{Prob}(y_i) = \frac{\Gamma(y_i + n_i)}{\Gamma(n_i) \Gamma(y_i + 1)} p_i^{n_i} (1 - p_i)^{y_i}, \quad y_i = 0, 1, 2, \ldots, \quad i = 1, 2, \dots, m\]
where:
\(n_i > 0\) is the number of successful trials (size) or dispersion parameter. Must be strictly positive but need not be an integer.
\(p_i \in (0, 1)\) is the probability of success in each trial.
Mean and Variance
The mean and variance of each observation are:
\[\mu_i = n_i \frac{1 - p_i}{p_i}, \qquad \sigma_i^2 = \mu_i \left(1 + \frac{\mu_i}{n_i}\right), \quad i = 1, 2, \dots, m\]
Note that the variance exceeds the mean (overdispersion), which distinguishes the negative binomial from the Poisson distribution.
Link Function
The mean is linked to the linear predictor \(\pmb{\eta} = (\eta_1, \eta_2, \dots, \eta_m)\) by:
\[\mu_i = E_i \exp(\eta_i), \quad i = 1, 2, \dots, m\]
or in vector form:
\[\pmb{\mu} = \pmb{E} \circ \exp(\pmb{\eta})\]
where \(\pmb{E} = (E_1, E_2, \dots, E_m)\) represents known constants (exposure), and \(\log(\pmb{E})\) is the offset of \(\pmb{\eta}\).
Possible link functions: default, log, logoffset.
Hyperparameters
The hyperparameter is the dispersion parameter \(n\) (size), which depends on the chosen variant:
variant=0 (default): The dispersion parameter is a scalar:
\[n = \exp(\theta)\]
variant=1: The dispersion parameter scales with exposure:
\[n_i = E_i \exp(\theta), \quad i = 1, 2, \dots, m\]
variant=2: The dispersion parameter scales with scale:
\[n_i = s_i \exp(\theta), \quad i = 1, 2, \dots, m\]
where \(s_i\) is the scale for each observation, and the prior is defined on \(\theta\).
Key: size
Default prior specification:
Prior: pc.mgamma
Parameters: [7]
Initial value: log(10) = 2.303
Fixed: False (estimated)
When translated into control['family']['hyper'], the default entry looks like this:
control = {
'family': {
'hyper': [{
'id': 'size',
'prior': 'pc.mgamma',
'param': [7],
'initial': 2.303,
'fixed': False,
}]
}
}
Each entry in control['family']['hyper'] may contain these keys:
id- Hyperparameter identifier (size). Can be omitted for the first (and only) hyperparameter.prior- Prior distribution nameparam- Prior parameters (list)initial- Initial value on log scalefixed- Whether to fix the hyperparameter (True/False)
Note: The PC-prior is available for variant=1.
Validation Rules
pyINLA enforces several validation rules for nbinomial models to ensure correct specification:
Exposure (E)
Key: E
When providing the E (exposure) argument:
All values must be strictly positive (> 0)
Length must match the number of observations
# With exposure
model = {"response": "y", "fixed": ["1", "x"]}
result = pyinla(model=model, family="nbinomial", data=df, E=df["exposure"].to_numpy())Scale
Key: scale
When providing the scale argument (used with variant=2):
All values must be strictly positive (> 0)
Length must match the number of observations
# With scale (variant=2)
model = {"response": "y", "fixed": ["1", "x"]}
result = pyinla(
model=model, family="nbinomial", data=df,
E=df["E"].to_numpy(),
scale=df["scale"].to_numpy(),
control={"family": {"variant": 2}}
)Variant
Key: control['family']['variant']
The variant must be one of:
0(default): Scalar dispersion parameter1: Dispersion scales with exposure2: Dispersion scales with scale
# variant=1: dispersion scales with exposure
model = {"response": "y", "fixed": ["1", "x"]}
result = pyinla(
model=model, family="nbinomial", data=df,
E=df["E"].to_numpy(),
control={"family": {"variant": 1}}
)Hyperparameters
Key: control['family']['hyper']
Hyperparameter configuration is allowed for nbinomial. The size hyperparameter controls the dispersion.
# Custom prior on dispersion parameter
model = {"response": "y", "fixed": ["1", "x"]}
control = {
'family': {
'hyper': [{
'id': 'size',
'prior': 'pc.mgamma',
'param': [10.0]
}]
}
}
result = pyinla(model=model, family="nbinomial", data=df, control=control)
# Fixed dispersion (not estimated)
control = {
'family': {
'hyper': [{
'id': 'size',
'initial': 2.303, # log(10), so size = 10
'fixed': True
}]
}
}
result = pyinla(model=model, family="nbinomial", data=df, control=control)
Ntrials Not Allowed
The Ntrials argument is not allowed for nbinomial. Use nbinomial2 if you need fixed trial counts.
Allowed Link Functions
Key: control['family']['link']
These link functions are supported:
log(default)logoffset
Response Values
Response variable \(\pmb{y}\) must be non-negative integers (counts: 0, 1, 2, ...). pyINLA will raise PyINLAError if any response value is negative or non-integer.
Specification
family="nbinomial"Required arguments: \(\pmb{y}\) (response), \(\pmb{E}\) (exposure, default = 1), and
scale(default = 1).Choose variant with
control={'family': {'variant': 0}}(default),{'variant': 1}, or{'variant': 2}.
Notes
As \(n \to \infty\), the negative binomial converges to the Poisson distribution. For numerical reasons, if \(n\) is too large:
\[\frac{\mu}{n} < 10^{-4}\]
then the Poisson limit is used.
The nbinomial2 Distribution
The negative binomial distribution is also available in its "pure form" as the number of excess experiments to get \(n\) successes with a success in the last experiment:
\[\text{Prob}(y_i) = \binom{y_i + n_i - 1}{n_i - 1} (1 - p_i)^{y_i} p_i^{n_i}, \quad y_i = 0, 1, 2, \ldots, \quad i = 1, 2, \dots, m\]
where:
\(n_i = 1, 2, \ldots\) is the (fixed) number of successes before stopping.
\(p_i\) is the probability of success in each independent trial.
Link Function for nbinomial2
The probability \(p_i\) is linked to the linear predictor \(\eta_i\) via the logit link (default):
\[p_i = \frac{\exp(\eta_i)}{1 + \exp(\eta_i)}, \quad i = 1, 2, \dots, m\]
Possible link functions: default, logit, loga, cauchit, probit, cloglog, ccloglog, loglog.
Hyperparameters for nbinomial2
None.
Validation Rules for nbinomial2
pyINLA enforces several validation rules for nbinomial2 models to ensure correct specification:
Ntrials (Required)
Key: Ntrials
The Ntrials argument is required for nbinomial2:
Must be provided for each observation
Values must be positive integers
Length must match the number of observations
# nbinomial2 requires Ntrials
model = {"response": "y", "fixed": ["1", "x"]}
result = pyinla(model=model, family="nbinomial2", data=df, Ntrials=df["n"].to_numpy())Exposure Not Allowed
The E (exposure) argument is not allowed for nbinomial2. Use nbinomial if you need exposure.
Scale Not Allowed
The scale argument is not allowed for nbinomial2. Use nbinomial if you need scale.
No Hyper Configuration
Key: control['family']['hyper']
The nbinomial2 likelihood has no hyperparameters. Do NOT provide control['family']['hyper'] configuration.
Allowed Link Functions
Key: control['family']['link']
These link functions are supported:
logit(default)logacauchitprobitcloglogccloglogloglog
Specification for nbinomial2
family="nbinomial2"Required arguments: \(\pmb{y}\) (response) and
Ntrials(the value of \(n\) for each observation).