Negative Binomial Regression
A tutorial on Bayesian Negative Binomial regression for overdispersed count data using pyINLA.
Introduction
In this tutorial, we fit a Bayesian Negative Binomial regression model for count data with overdispersion. The Negative Binomial distribution is the natural choice when modeling count data where the variance exceeds the mean, which is common in ecology, epidemiology, and many other fields.
Unlike the Poisson distribution (where variance equals the mean), the Negative Binomial has an additional dispersion parameter that captures extra-Poisson variation.
The Model
We consider a Negative Binomial regression model with $m$ observations. For each observation $i = 1, \ldots, m$, the count response $y_i$ follows:
where:
- $y_i \in \{0, 1, 2, \ldots\}$ is the count response
- $n > 0$ is the dispersion parameter (size)
- $p_i \in (0, 1)$ is the probability of success
The mean and variance are:
Note that $\sigma_i^2 > \mu_i$ (overdispersion), which distinguishes this from the Poisson distribution.
Linear Predictor
The linear predictor $\eta_i$ links to the mean through a log-link function:
where $\beta_0$ is the intercept and $\beta_1$ is the slope coefficient for covariate $x_i$.
The Negative Binomial distribution uses the log link function (default), so the mean is:
This ensures that the predicted mean count is always positive.
Likelihood Function
The likelihood for the full dataset $\mathbf{y} = (y_1, \ldots, y_m)^\top$ is:
where $\mu_i = \exp(\beta_0 + \beta_1 x_i)$ and $p_i = n / (n + \mu_i)$.
Hyperparameters
The Negative Binomial likelihood has one hyperparameter: the dispersion parameter $n$ (also called size). It is internally represented as:
where the prior is placed on $\theta$. The default prior is pc.mgamma with initial value 2.303.
As $n \to \infty$, the Negative Binomial converges to the Poisson distribution. Smaller values of $n$ indicate greater overdispersion.
Dataset
The dataset contains $m = 1000$ observations with two columns:
| Column | Description | Type |
|---|---|---|
y | Count response (non-negative integers) | int |
x | Covariate (predictor variable) | float |
Download the CSV file and place it in your working directory.
Implementation in pyINLA
We fit the model using pyINLA. The key steps are:
- Load the data into a pandas DataFrame
- Define the model structure (response and fixed effects)
- Call
pyinla()withfamily='nbinomial'
import pandas as pd
from pyinla import pyinla
# Load data
df = pd.read_csv('dataset_nbinomial_regression.csv')
# Define model: y ~ 1 + x (intercept + slope)
model = {
'response': 'y',
'fixed': ['1', 'x']
}
# Control settings (variant=0 is default)
control = {
'family': {'variant': 0}
}
# Fit the Negative Binomial regression model
result = pyinla(
model=model,
family='nbinomial',
data=df,
control=control
)
# View posterior summaries for fixed effects
print(result.summary_fixed)
# View posterior summary for dispersion parameter
print(result.summary_hyperpar)
Extracting Results
The result object contains all posterior inference:
# Fixed effects: posterior summaries for beta_0 and beta_1
print(result.summary_fixed)
# Extract posterior means
intercept = result.summary_fixed.loc['(Intercept)', 'mean']
slope = result.summary_fixed.loc['x', 'mean']
# Compute predicted means
import numpy as np
eta_hat = intercept + slope * df['x'].to_numpy()
mu_hat = np.exp(eta_hat)
# Hyperparameter: dispersion parameter n (size)
print(result.summary_hyperpar)
The summary_fixed DataFrame includes columns for mean, sd, 0.025quant, 0.5quant, 0.975quant, and mode.
Results and Diagnostics
The posterior summaries provide estimates for $\beta_0$, $\beta_1$, and the dispersion parameter $n$. The figures below show the model fit and residual diagnostics.
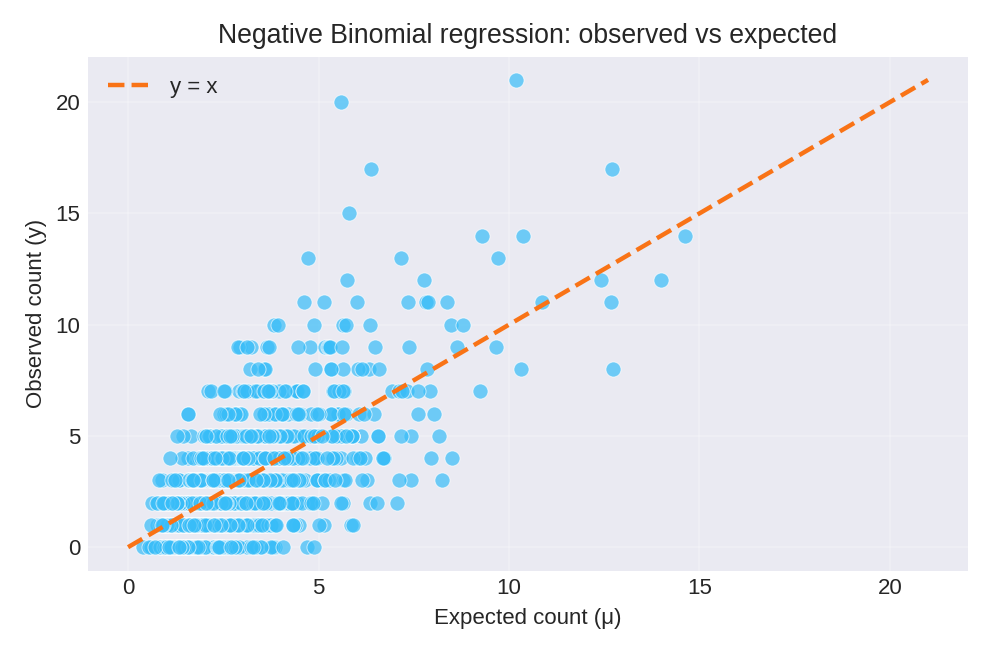
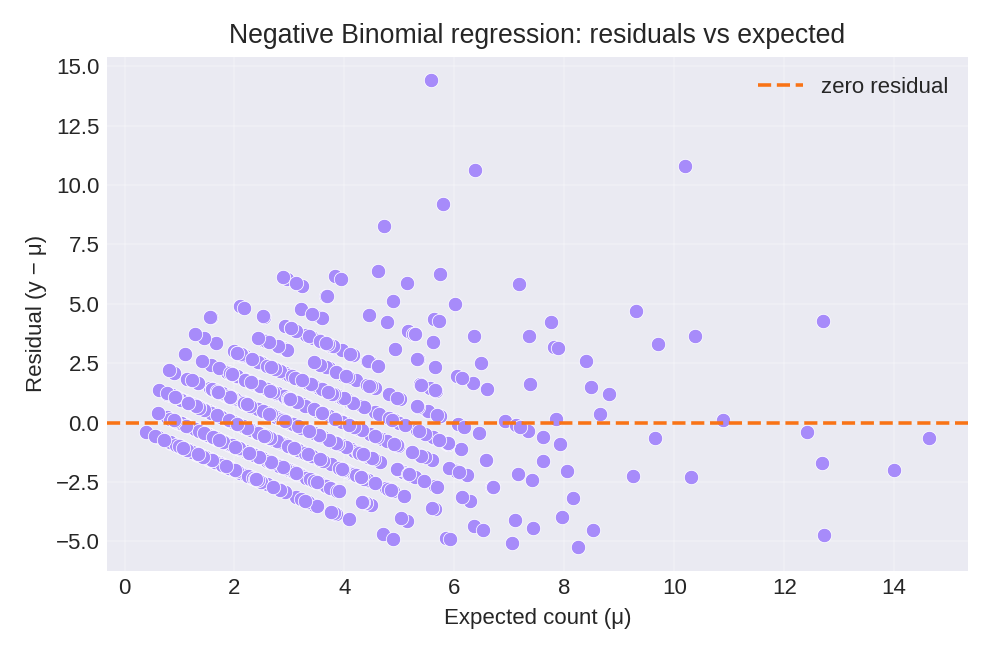
To reproduce these figures locally, download the render_nbinomial_plots.py script and run it alongside the CSV dataset.
Data Generation (Reference)
For reproducibility, the dataset was simulated from the following data-generating process:
where $\eta_i = \beta_0 + \beta_1 x_i$, $\mu_i = \exp(\eta_i)$, and $p_i = n / (n + \mu_i)$.
True parameter values: $\beta_0 = 1$, $\beta_1 = 0.5$, $n = 10$ (dispersion).
# Data generation script (for reference only)
import numpy as np
import pandas as pd
rng = np.random.default_rng(2026)
m = 1000
beta0, beta1 = 1.0, 0.5
size = 10.0 # dispersion parameter
x = rng.normal(size=m)
eta = beta0 + beta1 * x
mu = np.exp(eta)
# Negative binomial: p = size / (size + mu)
p = size / (size + mu)
y = rng.negative_binomial(n=size, p=p)
df = pd.DataFrame({'y': y, 'x': np.round(x, 6)})
# df.to_csv('dataset_nbinomial_regression.csv', index=False)