Parametrization
The Beta-Binomial distribution for a random vector \(\pmb{y} = (y_1, y_2, \dots, y_n)\) of count observations arises from a hierarchical model. The probability of success \(p_i\) follows a Beta distribution:
\[\pi(p_i) = \frac{1}{B(\alpha, \beta)} p_i^{\alpha-1} (1-p_i)^{\beta-1}, \quad \alpha > 0, \; \beta > 0, \quad i = 1, 2, \dots, n\]
Given \(p_i\), the response \(y_i\) is Binomial:
\[\pi(y_i \mid p_i) = \binom{N_i}{y_i} p_i^{y_i} (1-p_i)^{N_i-y_i}, \quad y_i = 0, 1, \ldots, N_i\]
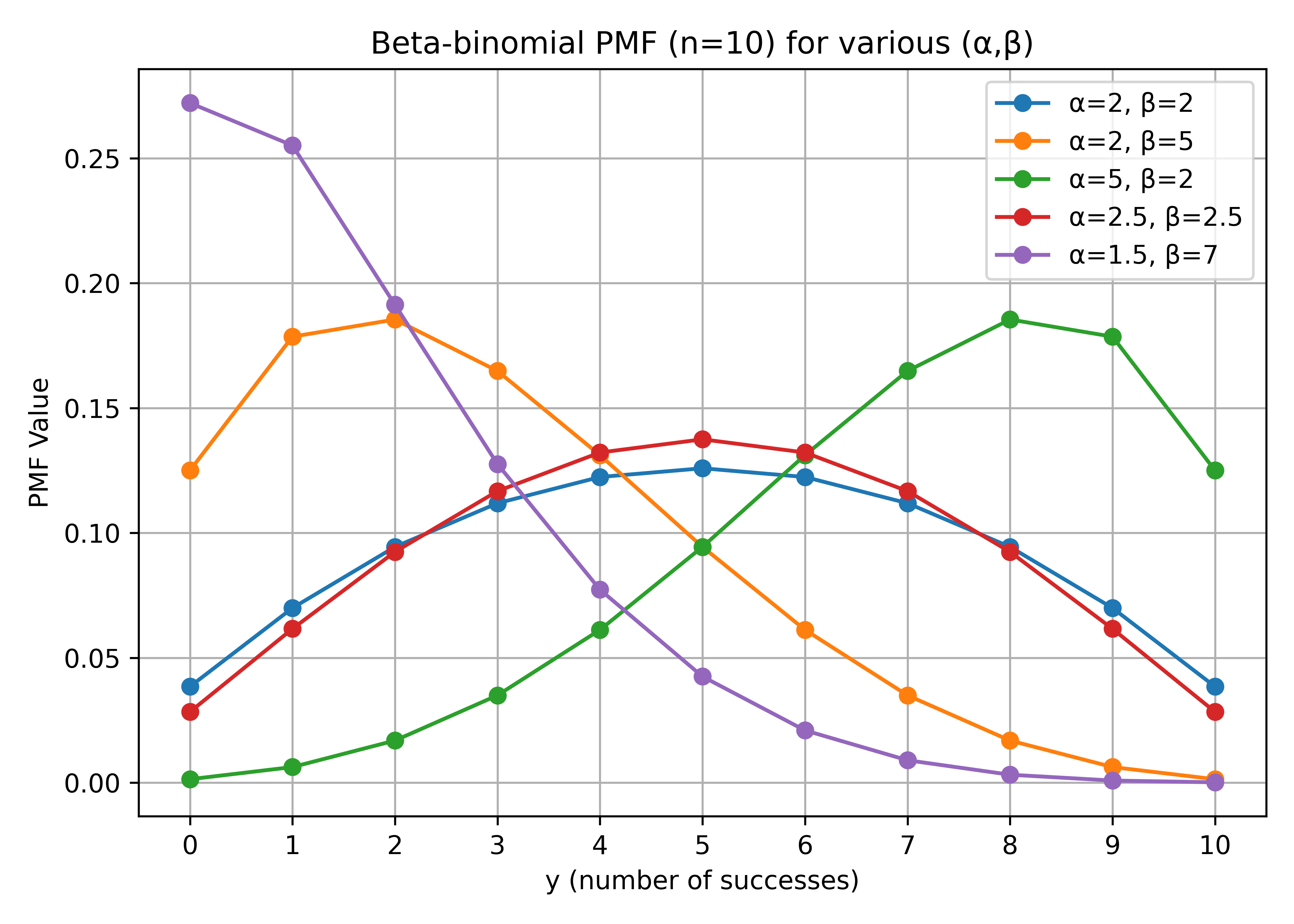
Marginalizing over \(p_i\), the distribution of \(y_i\) is Beta-Binomial:
\[\pi(y_i) = \binom{N_i}{y_i} \frac{B(y_i + \alpha, N_i - y_i + \beta)}{B(\alpha, \beta)}, \quad i = 1, 2, \dots, n\]
where \(B(\alpha, \beta) = \frac{\Gamma(\alpha)\Gamma(\beta)}{\Gamma(\alpha+\beta)}\) is the Beta function.
Mean and Variance
The mean and variance of each observation are:
\[\mu_i = N_i \mu_p, \qquad \sigma_i^2 = N_i \mu_p (1 - \mu_p) \left(1 + (N_i - 1) \rho\right), \quad i = 1, 2, \dots, n\]
where:
\(\mu_p = \frac{\alpha}{\alpha + \beta}\) is the mean probability from the Beta distribution.
\(\rho = \frac{1}{\alpha + \beta + 1}\) is the overdispersion parameter (\(0 < \rho < 1\)).
The term \((1 + (N_i - 1)\rho)\) inflates the variance beyond the standard Binomial, capturing overdispersion.
Link Function
The mean probability \(\mu_p\) is linked to the linear predictor \(\pmb{\eta} = (\eta_1, \eta_2, \dots, \eta_n)\) using the logit link (default):
\[\mu_p = \frac{\exp(\eta_i)}{1 + \exp(\eta_i)}, \quad i = 1, 2, \dots, n\]
Possible link functions: default, logit, loga, cauchit, probit, cloglog, ccloglog, loglog, robit, sn.
Hyperparameters
The hyperparameter is the overdispersion parameter \(\rho\), which is represented as:
\[\rho = \frac{\exp(\theta)}{1 + \exp(\theta)}\]
where the prior is defined on \(\theta\), ensuring \(\rho \in (0, 1)\).
Key: rho
Default prior specification:
Prior: gaussian
Parameters: (mean = 0, precision = 0.4)
Initial value: 0
Validation Rules
pyINLA enforces several validation rules for betabinomial models to ensure correct specification:
Ntrials (Required)
Key: Ntrials
The Ntrials argument is required for betabinomial:
Must be provided for each observation
Values must be positive integers
Length must match the number of observations
# betabinomial requires Ntrials
model = {"response": "y", "fixed": ["1", "x"]}
result = pyinla(model=model, family="betabinomial", data=df, Ntrials=df["n"].to_numpy())Hyperparameters
Key: control['family']['hyper']
Hyperparameter configuration is allowed for betabinomial. The rho hyperparameter controls the overdispersion.
Each entry in control['family']['hyper'] may contain these keys:
id- Hyperparameter identifier (rho). Can be omitted for the first (and only) hyperparameter.prior- Prior distribution nameparam- Prior parameters (list)initial- Initial value on logit scalefixed- Whether to fix the hyperparameter (True/False)
# Custom hyperparameter configuration (default values shown)
model = {"response": "y", "fixed": ["1", "x"]}
control = {
'family': {
'hyper': [{
'id': 'rho',
'prior': 'gaussian',
'param': [0.0, 0.4],
'initial': 0,
'fixed': False
}]
}
}
result = pyinla(model=model, family="betabinomial", data=df,
Ntrials=df["n"].to_numpy(), control=control)
# Fixed overdispersion (not estimated)
control = {
'family': {
'hyper': [{
'id': 'rho',
'initial': 0.0, # logit(0.5) = 0, so rho = 0.5
'fixed': True
}]
}
}
result = pyinla(model=model, family="betabinomial", data=df,
Ntrials=df["n"].to_numpy(), control=control)
Variant
Key: control['family']['variant']
The variant parameter is allowed for betabinomial:
0(default): Standard parameterization1: Alternative parameterization
Response Values
Response variable \(\pmb{y}\) must be non-negative integers with \(0 \leq y_i \leq N_i\) for each observation. pyINLA will raise PyINLAError if any response value is negative, non-integer, or exceeds its corresponding Ntrials value.
Exposure Not Allowed
The E (exposure) argument is not allowed for betabinomial. Use nbinomial or poisson if you need exposure.
Scale Not Allowed
The scale argument is not allowed for betabinomial.
Allowed Link Functions
Key: control['family']['link']
These link functions are supported:
logit(default)logacauchitprobitcloglogccloglogloglogrobitsn
# With probit link
model = {"response": "y", "fixed": ["1", "x"]}
result = pyinla(model=model, family="betabinomial", data=df,
Ntrials=df["n"].to_numpy(), control={"family": {"link": "probit"}})Specification
family="betabinomial"Required arguments: \(\pmb{y}\) (response) and
Ntrials(\(N_i\)).Optional:
control['family']['variant'](0 or 1),control['family']['hyper'],control['family']['link'].