Beta-Binomial Regression
A tutorial on Bayesian Beta-Binomial regression for overdispersed binomial data using pyINLA.
Introduction
In this tutorial, we fit a Bayesian Beta-Binomial regression model for count data arising from repeated Bernoulli trials with overdispersion. The Beta-Binomial distribution is the natural choice when the probability of success varies across observations, leading to extra-binomial variation.
Unlike the standard Binomial distribution (where success probability is fixed), the Beta-Binomial has an overdispersion parameter that captures heterogeneity in success probabilities across observations.
The Model
We consider a Beta-Binomial regression model with $n$ observations. For each observation $i = 1, \ldots, n$, the count response $y_i$ follows a hierarchical model:
Marginalizing over $p_i$, the response follows a Beta-Binomial distribution:
where $B(\cdot, \cdot)$ is the Beta function and $N_i$ is the number of trials for observation $i$.
Mean and Variance
The mean and variance of each observation are:
where:
- $\mu_p = \frac{\alpha}{\alpha + \beta}$ is the mean probability from the Beta distribution
- $\rho = \frac{1}{\alpha + \beta + 1}$ is the overdispersion parameter ($0 < \rho < 1$)
The term $(1 + (N_i - 1)\rho)$ inflates the variance beyond the standard Binomial, capturing overdispersion. As $\rho \to 0$, the distribution converges to Binomial.
Linear Predictor
The linear predictor $\eta_i$ links to the mean probability through the logit link function (default):
where $\beta_0$ is the intercept and $\beta_1$ is the slope coefficient for covariate $z_i$.
The mean probability is then:
This ensures that the predicted probability is always in $(0, 1)$.
Hyperparameters
The Beta-Binomial likelihood has one hyperparameter: the overdispersion parameter $\rho$. It is internally represented as:
where the prior is placed on $\theta$. The default prior is Gaussian(0, 0.4).
As $\rho \to 0$, the Beta-Binomial converges to the standard Binomial distribution. Larger values of $\rho$ indicate greater overdispersion.
Dataset
The dataset contains $n = 1000$ observations with three columns:
| Column | Description | Type |
|---|---|---|
y | Count response (successes) | int |
z | Covariate (predictor variable) | float |
Ntrials | Number of trials per observation | int |
Download the CSV file and place it in your working directory.
Implementation in pyINLA
We fit the model using pyINLA. The key steps are:
- Load the data into a pandas DataFrame
- Define the model structure (response and fixed effects)
- Call
pyinla()withfamily='betabinomial'and passNtrials
import pandas as pd
from pyinla import pyinla
# Load data
df = pd.read_csv('dataset_betabinomial_regression.csv')
Ntrials = df['Ntrials'].to_numpy()
# Define model: y ~ 1 + z (intercept + slope)
model = {
'response': 'y',
'fixed': ['1', 'z']
}
# Fit the Beta-Binomial regression model
result = pyinla(
model=model,
family='betabinomial',
data=df,
Ntrials=Ntrials
)
# View posterior summaries for fixed effects
print(result.summary_fixed)
# View posterior summary for overdispersion parameter
print(result.summary_hyperpar)
Extracting Results
The result object contains all posterior inference:
# Fixed effects: posterior summaries for beta_0 and beta_1
print(result.summary_fixed)
# Extract posterior means
intercept = result.summary_fixed.loc['(Intercept)', 'mean']
slope = result.summary_fixed.loc['z', 'mean']
# Compute predicted probabilities
import numpy as np
eta_hat = intercept + slope * df['z'].to_numpy()
p_hat = np.exp(eta_hat) / (1 + np.exp(eta_hat))
# Hyperparameter: overdispersion parameter rho
print(result.summary_hyperpar)
The summary_fixed DataFrame includes columns for mean, sd, 0.025quant, 0.5quant, 0.975quant, and mode.
Data Generation (Reference)
For reproducibility, the dataset was simulated from the following data-generating process:
With overdispersion $\rho = 0.2$ and linear predictor $\eta_i = 0.5 + z_i$:
Then $p_i \sim \text{Beta}(a_i, b_i)$ and $y_i \sim \text{Binomial}(N_i, p_i)$.
True parameter values: $\beta_0 = 0.5$, $\beta_1 = 1.0$, $\rho = 0.2$.
# Data generation script (for reference only)
import numpy as np
import pandas as pd
rng = np.random.default_rng(2026)
n = 1000
rho = 0.2 # moderate overdispersion
# Generate covariate and Ntrials
z = rng.normal(0, 1, size=n)
Ntrials = rng.integers(5, 21, size=n) # 5 to 20 inclusive
# Linear predictor and mean probability
beta0, beta1 = 0.5, 1.0
eta = beta0 + beta1 * z
p_eta = np.exp(eta) / (1 + np.exp(eta))
# Beta parameters from rho and p_eta
a = p_eta * (1 - rho) / rho
b = (1 - p_eta) * (1 - rho) / rho
# Generate probability from Beta, then response from Binomial
p = rng.beta(a, b)
y = rng.binomial(Ntrials, p)
df = pd.DataFrame({'y': y, 'z': np.round(z, 6), 'Ntrials': Ntrials})
# df.to_csv('dataset_betabinomial_regression.csv', index=False)
Results and Diagnostics
The posterior summaries recover the true parameters: $\beta_0 = 0.5$, $\beta_1 = 1.0$, and overdispersion $\rho = 0.2$. The diagnostic plots below compare observed counts with expected counts and show the residual pattern.
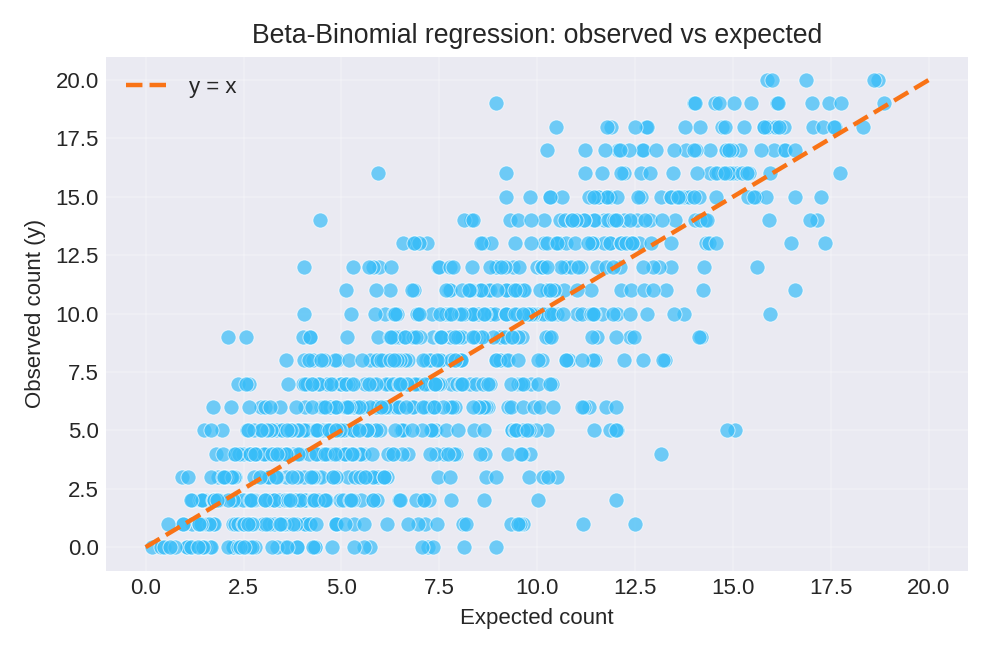
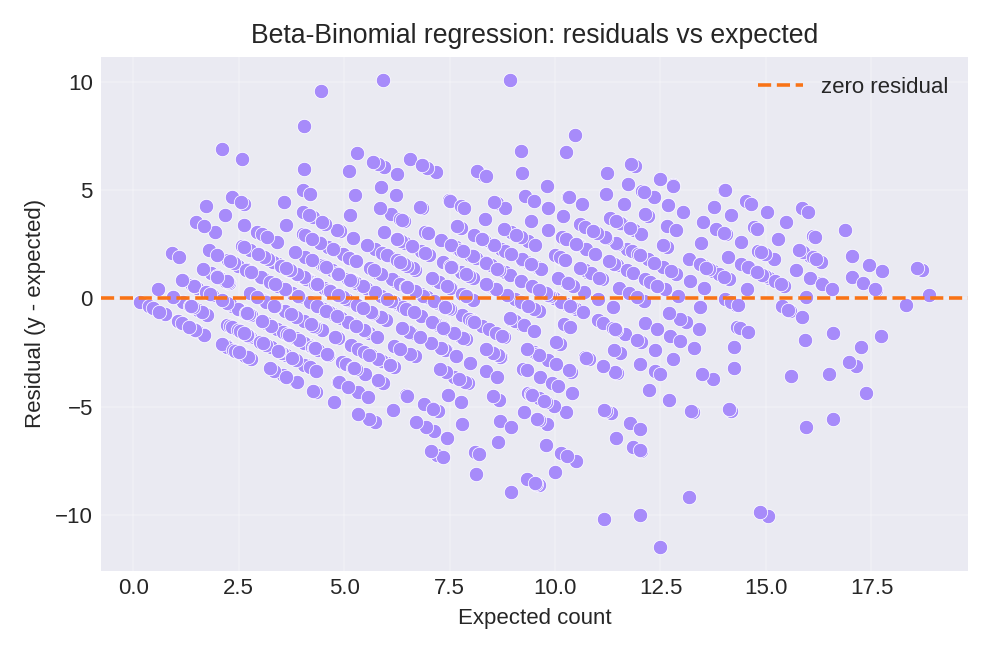
To reproduce these figures locally, download the render_betabinomial_plots.py script and run it alongside the CSV dataset.