The exponential distribution is a continuous probability distribution often used to model the time between independent events that occur at a constant average rate. It is characterized by its memoryless property, meaning the probability of an event occurring in the future is independent of the past. This distribution is widely utilized in reliability analysis and queuing theory.
Parametrization
The probability density function (PDF) for the exponential distribution, considering \(\pmb{y}\) as a vector of responses, is given by:
\[f(y_i \mid \lambda_i) = \lambda_i \exp(-\lambda_i y_i), \quad y_i > 0, \; \lambda_i > 0, \; i = 1, 2, \dots, n,\]
where:
\(\pmb{y} = (y_1, y_2, \ldots, y_n)\) represents the observed response times.
\(\pmb{\lambda} = (\lambda_1, \lambda_2, \ldots, \lambda_n)\) represents the rate parameters for each observation.
In survival analysis, models are often specified through the hazard function. For the exponential model, the baseline hazard is constant over time, and the hazard function is:
\[h(y_i) = \lambda_i, \quad i = 1, 2, \ldots, n. \\ \]
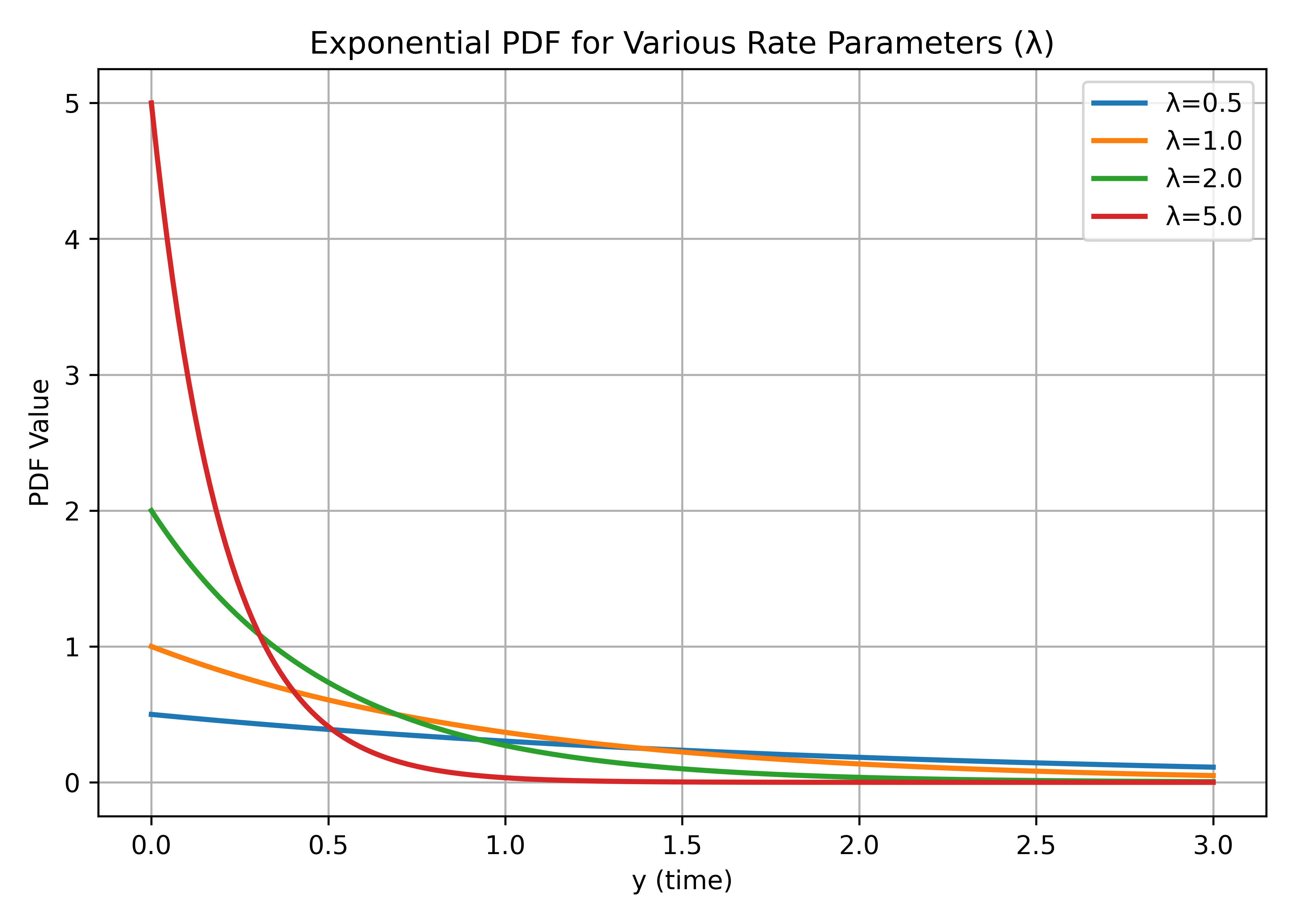
Assume \(n\) = 1. To demonstrate the shape of the Exponential distribution for various rate parameters \(\lambda\), Figure 1 displays several Exponential PDFs.
Link Function
The rate parameter \(\lambda_i\) is linked to the linear predictor \(\eta_i\) using the default log link:
\[\lambda_i = \exp(\eta_i), \quad i = 1, 2, \ldots, n.\]
The available link functions are: default, log.
Hyperparameters
The Exponential likelihood has no hyperparameters. The rate parameter \(\lambda\) is fully determined by the linear predictor \(\eta\) through the link function.
Validation Rules
pyINLA enforces several validation rules for Exponential models to ensure correct specification:
No Hyper Configuration
The Exponential likelihood has no hyperparameters. Do NOT provide control['family']['hyper'] configuration.
Not Allowed Arguments
The following arguments are not allowed for exponential likelihoods and will raise PyINLAError:
E(exposure) - Only allowed for poisson/nbinomialscale- Only allowed for gaussian/nbinomial/gamma/beta/logistic/tNtrials- Only allowed for binomial/betabinomial/nbinomial2control['family']['variant']- Not supported for exponential
Allowed Link Functions
Key: control['family']['link']
defaultlog
Response Values
Response variable \(\pmb{y}\) must be strictly positive (\(y_i > 0\)). pyINLA will raise PyINLAError if any response value is zero or negative.
Specification
family="exponential"- Required arguments:
- \(\pmb{y}\): response vector of strictly positive values