Gaussian Regression: Log-Gamma Prior
A complete tutorial on Bayesian linear regression with Gaussian likelihood and a log-gamma prior on the observation precision.
Introduction
In this tutorial, we fit a Bayesian linear regression model where the response variable follows a Gaussian (normal) distribution. The goal is to estimate the regression coefficients (intercept and slope) while treating the observation precision as an unknown hyperparameter with a log-gamma prior.
This example demonstrates the fundamental workflow in pyINLA: specifying a likelihood, defining the linear predictor, assigning priors to hyperparameters, and obtaining posterior inference.
The Model
We consider a simple linear regression model with $n$ observations. For each observation $i = 1, \ldots, n$, the response $y_i$ is modeled as:
where:
- $y_i \in \mathbb{R}$ is the continuous response variable
- $z_i \in \mathbb{R}$ is the covariate
- $\beta_0$ is the intercept
- $\beta_1$ is the slope coefficient
- $\tau > 0$ is the observation precision (inverse variance)
- $\varepsilon_i$ is the Gaussian noise term
Linear Predictor
The linear predictor $\eta_i$ links the mean of the response to the covariates through a linear combination:
For the Gaussian likelihood with identity link, the mean of the response equals the linear predictor:
In matrix notation, we can write $\boldsymbol{\eta} = \mathbf{X}\boldsymbol{\beta}$, where $\mathbf{X}$ is the $n \times 2$ design matrix with a column of ones (for the intercept) and a column containing the covariate values.
Likelihood Function
Using the linear predictor, we can write the model in terms of the conditional distribution of $y_i$:
The likelihood for the full dataset $\mathbf{y} = (y_1, \ldots, y_n)^\top$ is:
where $\boldsymbol{\beta} = (\beta_0, \beta_1)^\top$ denotes the vector of regression coefficients.
Prior on the Precision
In Bayesian inference, we need to specify a prior distribution for the unknown precision parameter $\tau$. A common choice is the log-gamma prior on the log-transformed precision $\theta = \log(\tau)$:
This is equivalent to placing a Gamma$(a, b)$ prior on $\tau$ itself:
The log-gamma parameterization is convenient for numerical optimization since $\theta \in \mathbb{R}$ is unconstrained.
Parameters used in this example: $a = 1.0$, $b = 0.01$, with initial value $\theta_0 = 2.0$ (corresponding to $\tau_0 \approx 7.4$).
Dataset
The dataset contains $n = 100$ observations with two columns:
| Column | Description | Type |
|---|---|---|
y | Continuous response variable | float |
z | Covariate (predictor variable) | float |
Download the CSV file and place it in your working directory.
Implementation in pyINLA
We now fit the model using pyINLA. The key steps are:
- Load the data into a pandas DataFrame
- Define the model structure (response and fixed effects)
- Specify the prior on the precision hyperparameter
- Call the
pyinla()function
import pandas as pd
from pyinla import pyinla
# Load data
df = pd.read_csv('dataset_gaussian_loggamma.csv')
# Define model: y ~ 1 + z (intercept + slope)
model = {
'response': 'y',
'fixed': ['1', 'z']
}
# Specify log-gamma prior on precision: Gamma(a=1.0, b=0.01)
control = {
'family': {
'hyper': [{
'prior': 'loggamma',
'param': [1.0, 0.01],
'initial': 2.0
}]
},
'predictor': {'compute': True} # enables fitted values
}
# Fit the model
result = pyinla(model=model, family='gaussian', data=df, control=control)
# View posterior summaries for fixed effects
print(result.summary_fixed)
Extracting Results
The result object contains all posterior inference. Here are the key attributes:
# Fixed effects: posterior summaries for beta_0 and beta_1
print(result.summary_fixed)
# Extract posterior means
intercept = result.summary_fixed.loc['(Intercept)', 'mean']
slope = result.summary_fixed.loc['z', 'mean']
# Fitted values: posterior mean of eta_i (requires predictor.compute = True)
fitted_means = result.summary_fitted_values['mean'].to_numpy()
# Compute residuals
residuals = df['y'].to_numpy() - fitted_means
# Hyperparameters: posterior summary for precision tau
print(result.summary_hyperpar)
# Marginal posterior density for tau (for plotting)
density = result.marginals_hyperpar['Precision for the Gaussian observations']
The summary_fixed DataFrame includes columns for mean, sd, 0.025quant, 0.5quant, 0.975quant, and mode.
Results and Diagnostics
The posterior summaries provide point estimates (mean, mode) and credible intervals for the regression coefficients $\beta_0$ and $\beta_1$, as well as the precision $\tau$.
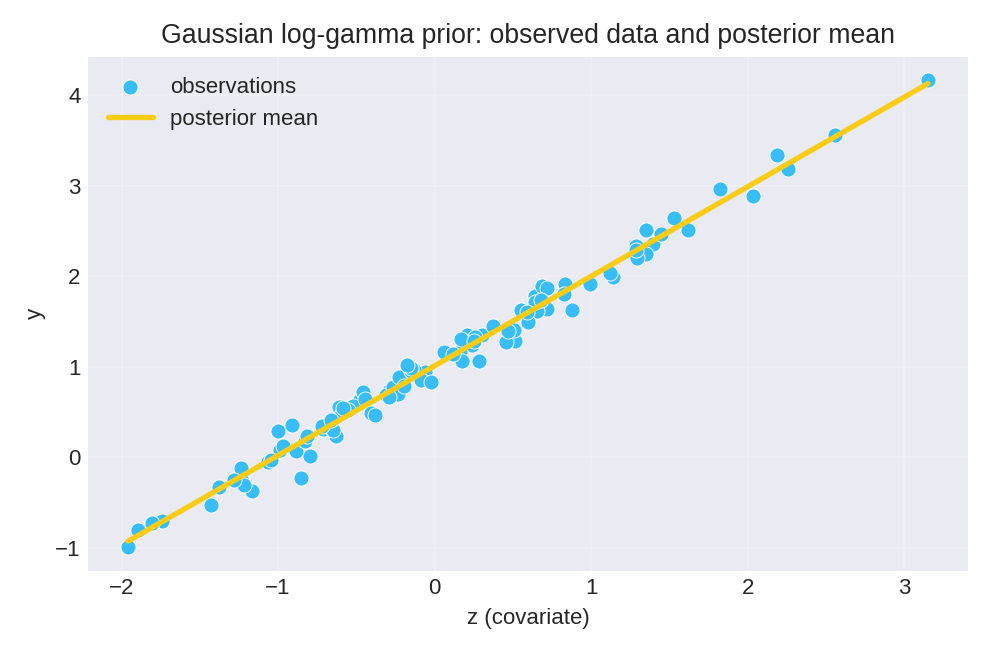
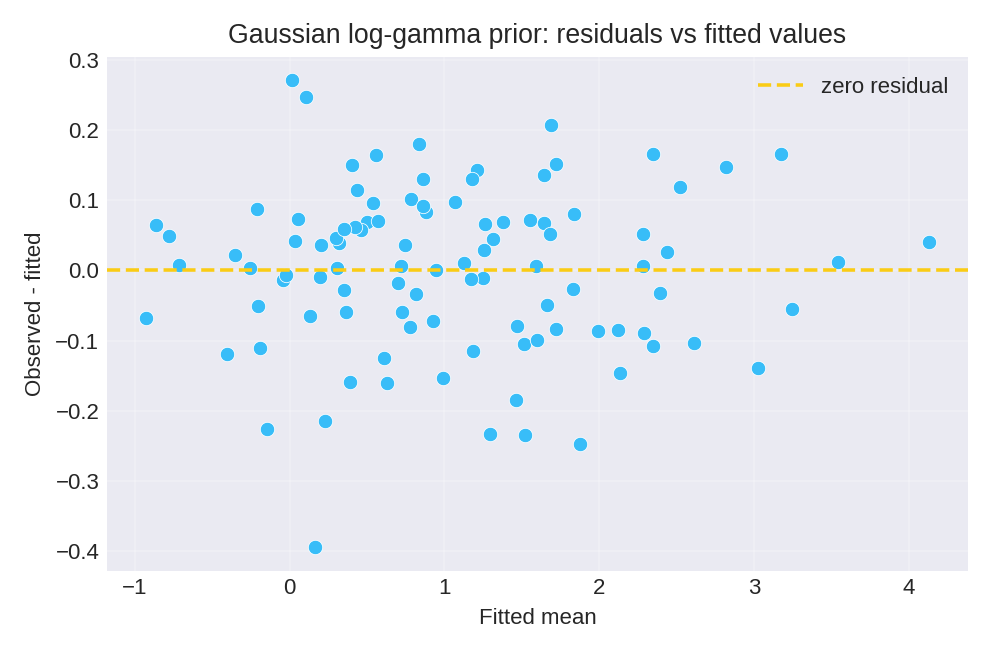
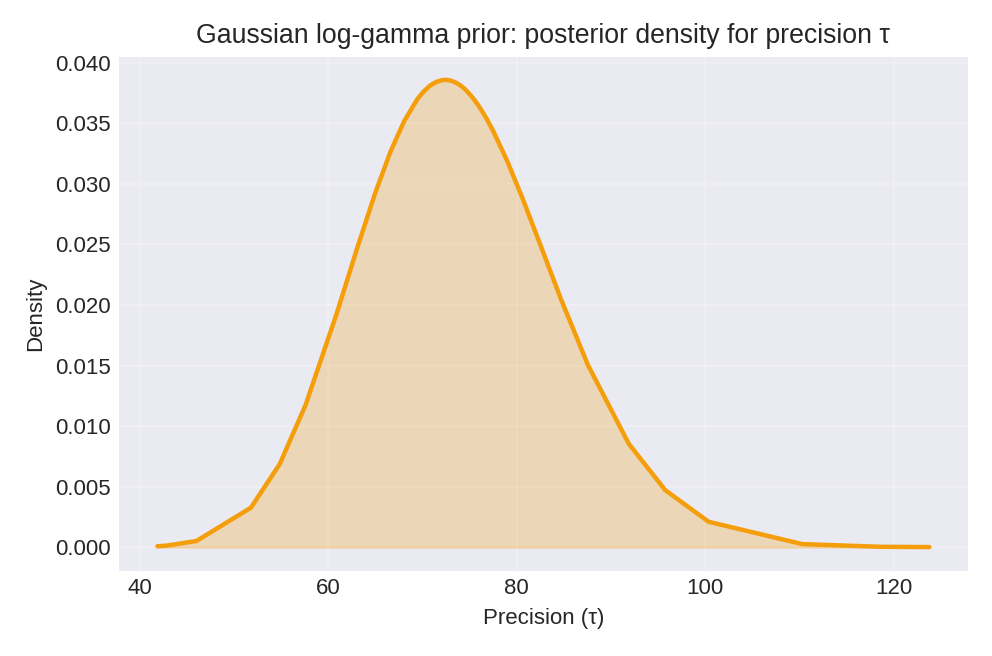
To reproduce these figures locally, download the render_gaussian_plots.py script and run it alongside the CSV dataset.
Data Generation (Reference)
For reproducibility, the dataset was simulated from the following data-generating process:
with true parameter values: $\beta_0 = 1$, $\beta_1 = 1$, $\tau = 100$ (i.e., $\sigma = 0.1$).
# Data generation script (for reference only)
import numpy as np
import pandas as pd
rng = np.random.default_rng(2026)
n = 100
beta0, beta1 = 1.0, 1.0
tau = 100.0 # precision (variance = 0.01)
z = rng.normal(size=n)
epsilon = rng.normal(scale=1.0/np.sqrt(tau), size=n)
y = beta0 + beta1 * z + epsilon
df = pd.DataFrame({'y': y, 'z': z})
# df.to_csv('dataset_gaussian_loggamma.csv', index=False)