Expert Binomial with Scale
Using the xbinomial family with a scale parameter to model probability-scaled binomial data.
Introduction
In this tutorial, we demonstrate the expert version of the binomial distribution (xbinomial) with a scale parameter. This is useful when the success probability needs to be scaled by a known factor for each observation.
The expert version also allows non-integer values for both $\pmb{y}$ and $\pmb{n}$, providing more flexibility for specialized modeling scenarios.
The Model
We consider an expert binomial regression model with $n$ observations. For each observation $i = 1, \ldots, n$, the count of successes $y_i$ out of $N_i$ trials is modeled as:
where the effective probability $p'_i$ is scaled:
and:
- $y_i \in \{0, 1, 2, \ldots, N_i\}$ is the observed number of successes
- $N_i > 0$ is the known number of trials for observation $i$
- $q_i \in (0, 1]$ is the known scale factor for observation $i$
- $p_i(\eta_i) = \text{logit}^{-1}(\eta_i)$ is the base probability from the linear predictor
Note: The "fitted values" reported by pyINLA will be $p(\eta)$, not $p' = q \cdot p(\eta)$.
Linear Predictor
The linear predictor $\eta_i$ links the log-odds of the base probability to the covariates:
where $\beta_0$ is the intercept and $\beta_1$ is the slope coefficient for covariate $x_i$.
The base probability is:
And the effective probability used in the binomial likelihood is:
Hyperparameters
Like the standard binomial, the expert binomial has no hyperparameters. The variance is determined by the mean:
Dataset
The dataset contains $n = 10,000$ observations with four columns:
| Column | Description | Type |
|---|---|---|
y | Number of successes | integer |
x | Covariate (predictor variable) | float |
q | Scale factor ($0 < q \le 1$) | float |
ntrials | Number of trials | integer |
Download the CSV file and place it in your working directory.
Implementation in pyINLA
We fit the model using pyINLA with family='xbinomial' and the scale parameter:
- Load the data into a pandas DataFrame
- Extract the ntrials and scale (q) vectors
- Define the model structure (response and fixed effects)
- Call
pyinla()withfamily='xbinomial',Ntrials, andscale
import pandas as pd
from pyinla import pyinla
# Load data
df = pd.read_csv('dataset_xbinomial.csv')
ntrials = df['ntrials'].to_numpy()
q = df['q'].to_numpy()
# Define model: y ~ 1 + x (intercept + slope)
model = {
'response': 'y',
'fixed': ['1', 'x']
}
# Fit the expert binomial model with scale
result = pyinla(
model=model,
family='xbinomial',
data=df,
Ntrials=ntrials,
scale=q
)
# View posterior summaries for fixed effects
print(result.summary_fixed)
Extracting Results
The result object contains all posterior inference:
# Fixed effects: posterior summaries for beta_0 and beta_1
print(result.summary_fixed)
# Extract posterior means
intercept = result.summary_fixed.loc['(Intercept)', 'mean']
slope = result.summary_fixed.loc['x', 'mean']
# Compute predicted base probabilities (not scaled)
import numpy as np
eta_hat = intercept + slope * df['x'].to_numpy()
p_hat = 1.0 / (1 + np.exp(-eta_hat))
# Compute effective probabilities (with scale)
p_effective = df['q'].to_numpy() * p_hat
# Compute expected counts
expected_counts = df['ntrials'].to_numpy() * p_effective
The summary_fixed DataFrame includes columns for mean, sd, 0.025quant, 0.5quant, 0.975quant, and mode.
Results and Diagnostics
The posterior summaries provide estimates for $\beta_0$ and $\beta_1$. The true parameters used for data generation were $\beta_0 = 0.88$ and $\beta_1 = 0.77$.
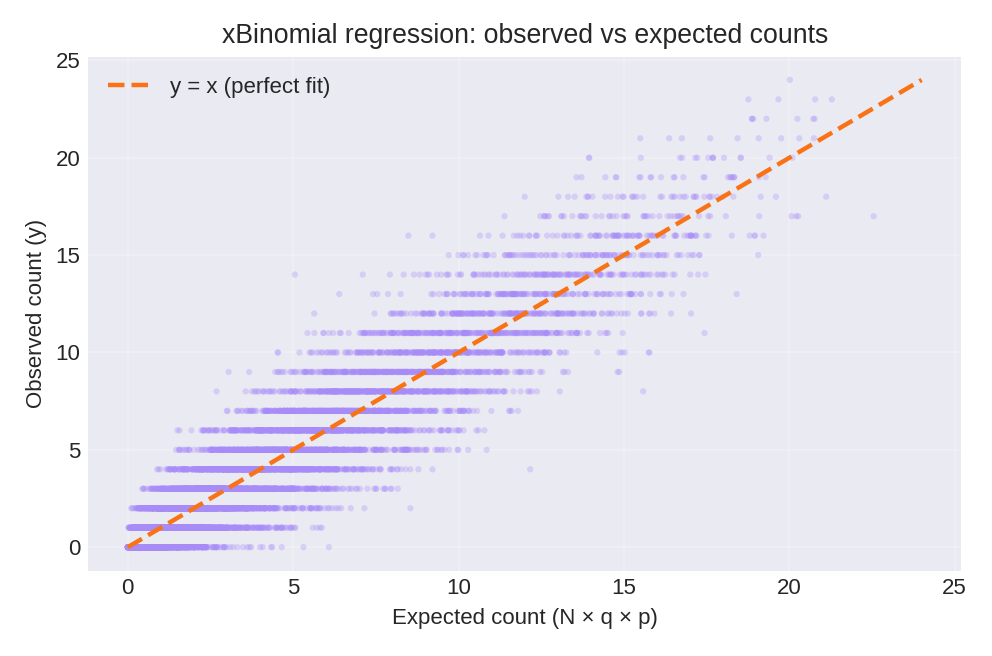
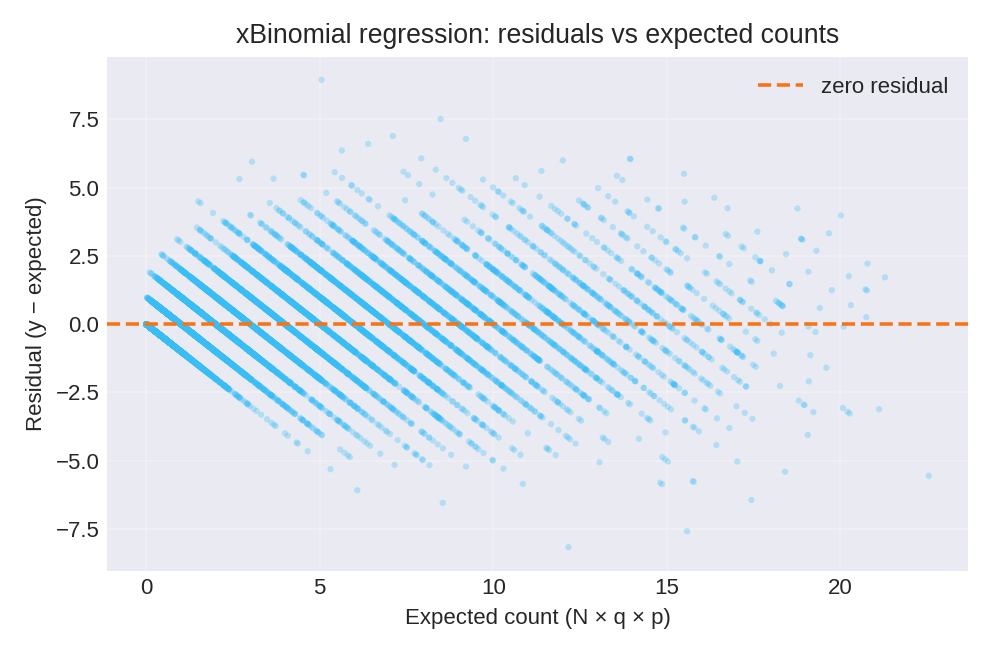
To reproduce these figures locally, download the render_xbinomial_plots.py script and run it alongside the CSV dataset.
Data Generation (Reference)
For reproducibility, the dataset was simulated from the following data-generating process:
where $\eta_i = \beta_0 + \beta_1 x_i$ with true parameter values: $\beta_0 = 0.88$, $\beta_1 = 0.77$.
# Data generation script (for reference only)
import numpy as np
import pandas as pd
rng = np.random.default_rng(123)
n = 10000
beta0, beta1 = 0.88, 0.77
x = rng.normal(size=n)
q = rng.uniform(0, 1, size=n) # scale factors
eta = beta0 + beta1 * x
p = 1.0 / (1 + np.exp(-eta))
p_scaled = q * p
ntrials = rng.integers(1, 26, size=n) # trials between 1 and 25
y = rng.binomial(n=ntrials, p=p_scaled)
df = pd.DataFrame({
'y': y,
'x': np.round(x, 6),
'q': np.round(q, 6),
'ntrials': ntrials
})
# df.to_csv('dataset_xbinomial.csv', index=False)