Beta Regression
A tutorial on Bayesian Beta regression for modeling proportions and rates using pyINLA.
Introduction
In this tutorial, we fit a Bayesian Beta regression model for data on the interval (0, 1). The Beta distribution is the natural choice when modeling proportions, rates, or probabilities where the response is continuous and bounded between 0 and 1.
The Beta likelihood uses a logit link function to ensure predictions stay within (0, 1), and includes a precision hyperparameter that controls the dispersion of the data around the mean.
The Model
We consider a Beta regression model with $n$ observations. For each observation $i = 1, \ldots, n$, the response $y_i \in (0, 1)$ is modeled as:
where the shape parameters $a_i$ and $b_i$ are related to the mean $\mu_i$ and precision $\phi_i$ through:
The mean and variance are:
Linear Predictor
The linear predictor $\eta_i$ links to the mean through a logit link function:
The Beta distribution uses the logit link function (default):
This ensures that the predicted mean is always in (0, 1).
Hyperparameters
The Beta likelihood has one hyperparameter: the precision parameter $\phi$. With a scale factor $s_i$:
where the prior is placed on $\theta$. The default prior is loggamma with parameters (1, 0.1).
Larger $\phi$ corresponds to smaller variance (more precision) for fixed $\mu$.
Dataset
The dataset contains $n = 1000$ observations with three columns:
| Column | Description | Type |
|---|---|---|
y | Response in (0, 1) | float |
z | Covariate (predictor variable) | float |
w | Scale factor for precision | float |
Download the CSV file and place it in your working directory.
Implementation in pyINLA
We fit the model using pyINLA with a varying precision through the scale parameter:
import pandas as pd
from pyinla import pyinla
# Load data
df = pd.read_csv('dataset_beta_regression.csv')
scale = df['w'].to_numpy()
# Define model: y ~ 1 + z (intercept + slope)
model = {
'response': 'y',
'fixed': ['1', 'z']
}
# Fit the Beta regression model
result = pyinla(
model=model,
family='beta',
data=df,
scale=scale
)
# View posterior summaries
print(result.summary_fixed)
print(result.summary_hyperpar)
Results and Diagnostics
The posterior summaries provide estimates for $\beta_0$ and the precision hyperparameter $\theta$. The figures below show the model fit and residual diagnostics.
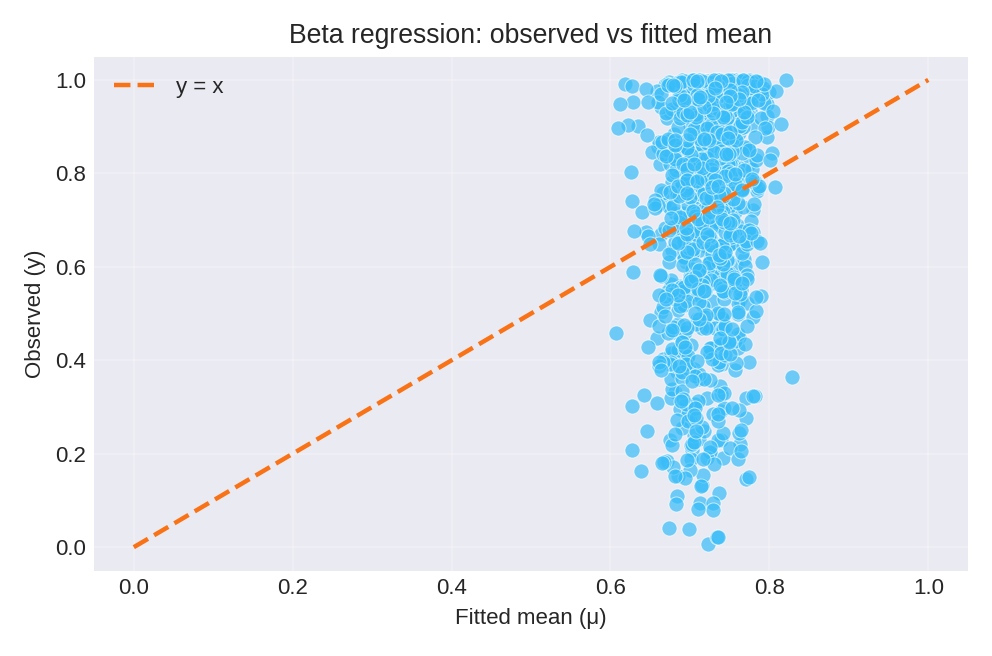
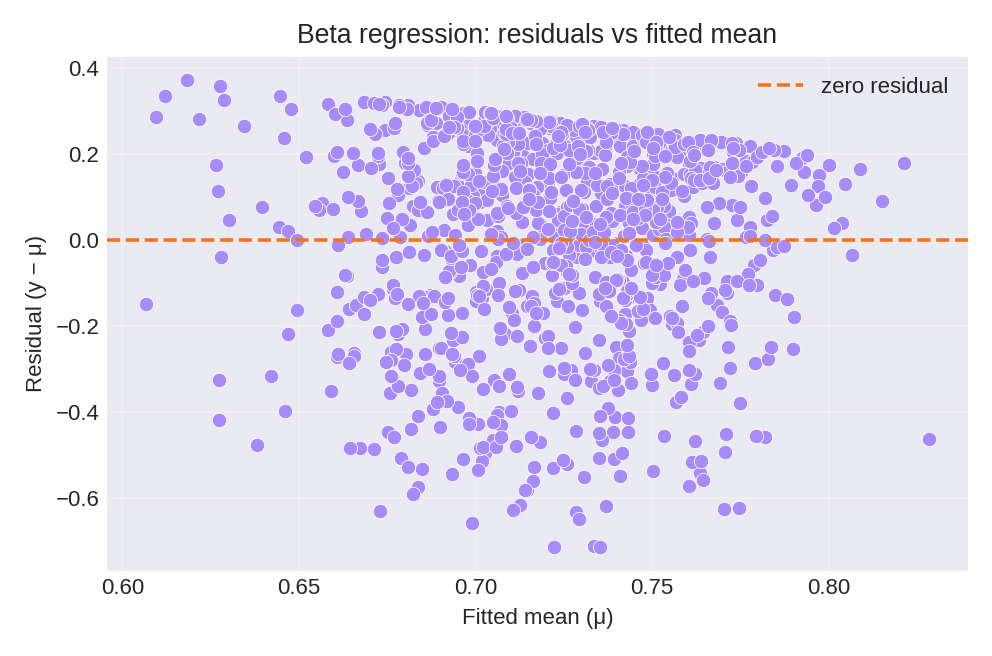
To reproduce these figures locally, download the render_beta_plots.py script and run it alongside the CSV dataset.
Data Generation (Reference)
For reproducibility, the dataset was simulated from:
with $\eta_i = 1 + z_i$, $\mu_i = \text{logit}^{-1}(\eta_i)$, $a_i = \mu_i \phi_i$, $b_i = \phi_i(1-\mu_i)$, and $y_i \sim \text{Beta}(a_i, b_i)$.
True parameter values: $\beta_0 = 1$, $\exp(\theta) = 5$.