Beta Regression with Censoring
Bayesian Beta regression with censored observations at the boundaries (0 and 1).
Introduction
In some applications, observations very close to 0 or 1 are recorded as exactly 0 or 1 due to measurement limitations or rounding. The Beta distribution is defined on the open interval (0, 1), so exact 0s and 1s require special handling.
The Beta likelihood in pyINLA supports censoring: observations at or beyond a threshold are treated as censored rather than exact values.
Censoring Mechanism
A censoring threshold $\delta \in (0, 0.5)$ is specified:
- Observations $y_i \leq \delta$ are treated as left-censored at 0
- Observations $y_i \geq 1 - \delta$ are treated as right-censored at 1
For example, with $\delta = 0.05$:
Dataset
The dataset contains $n = 1000$ observations with two columns:
| Column | Description | Type |
|---|---|---|
y | Response (may include 0s and 1s) | float |
z | Covariate (predictor variable) | float |
Download the CSV file and place it in your working directory.
Implementation in pyINLA
We fit the model using pyINLA with censoring enabled via beta.censor.value:
import pandas as pd
from pyinla import pyinla
# Load data
df = pd.read_csv('dataset_beta_censored.csv')
cens = 0.05 # censoring threshold
# Define model
model = {
'response': 'y',
'fixed': ['1', 'z']
}
# Control settings with censoring
control = {
'family': {
'beta.censor.value': cens
}
}
# Fit the Beta regression model with censoring
result = pyinla(
model=model,
family='beta',
data=df,
control=control
)
# View posterior summaries
print(result.summary_fixed)
print(result.summary_hyperpar)
Extracting Results
The result object contains all posterior inference:
import numpy as np
# Fixed effects: posterior summaries for beta_0 and beta_1
print(result.summary_fixed)
# Extract posterior means
intercept = result.summary_fixed.loc['(Intercept)', 'mean']
slope = result.summary_fixed.loc['z', 'mean']
# Compute predicted probabilities (logit link)
eta_hat = intercept + slope * df['z'].to_numpy()
mu_hat = np.exp(eta_hat) / (1 + np.exp(eta_hat))
# Compute residuals
residuals = df['y'].to_numpy() - mu_hat
# Hyperparameters: precision phi
print(result.summary_hyperpar)
The summary_fixed DataFrame includes columns for mean, sd, 0.025quant, 0.5quant, 0.975quant, and mode.
Results and Diagnostics
The posterior summaries provide estimates for $\beta_0$, $\beta_1$, and the precision $\phi$. Censored observations (at 0 and 1) are shown separately.
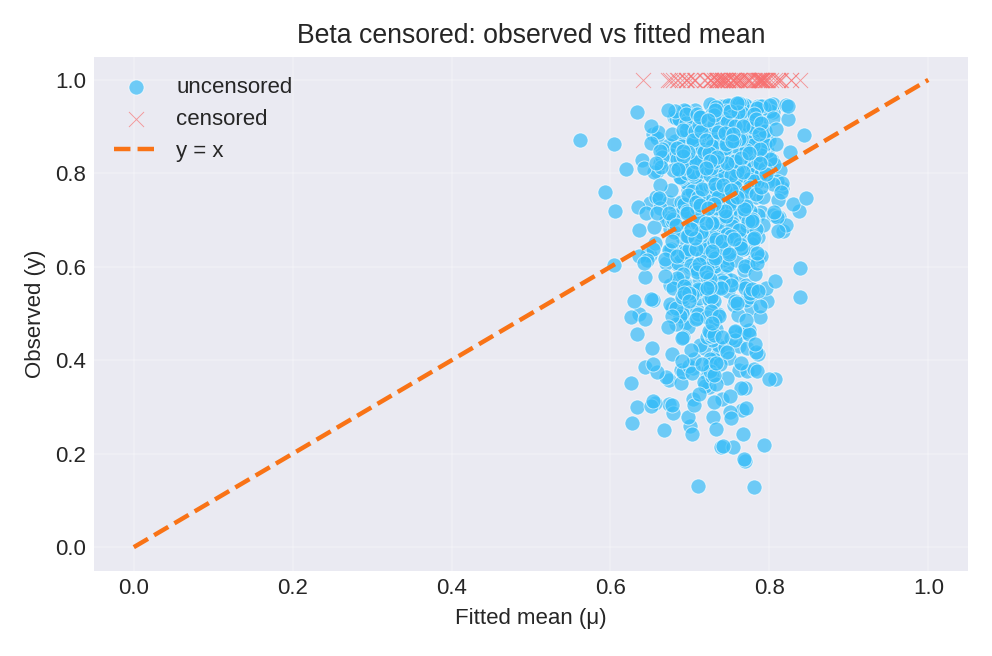
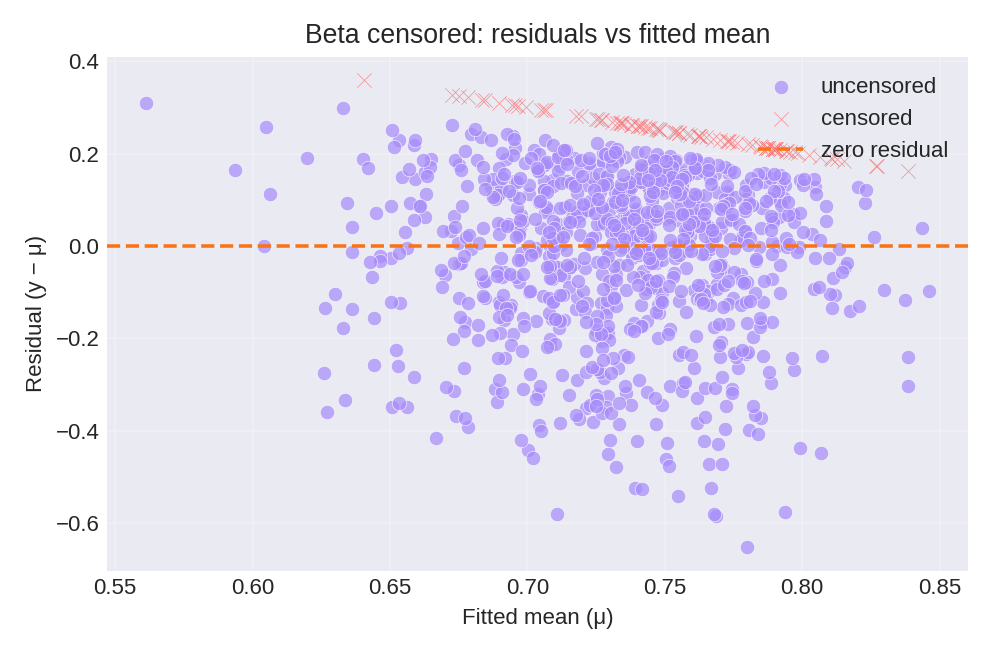
To reproduce these figures locally, download the render_beta_plots.py script and run it alongside the CSV dataset.
Data Generation (Reference)
For reproducibility, the dataset was simulated with fixed precision $\phi = 5$:
Then $y_i \sim \text{Beta}(\mu_i \phi, \phi(1-\mu_i))$ with censoring applied at $\delta = 0.05$.
True parameter values: $\beta_0 = 1$, $\phi = 5$.